Every Breath You Take
Shortening Treatment for Lung Cancer Patients
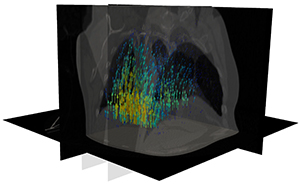 New research at the Scientific Computing and Imaging Institute at the University of Utah, helps to precisely target tumors in lung cancer patients using artificial intelligence. Employing algorithms developed by Sarang Joshi, DSc, professor of Biomedical Engineering and graduate student Markus Foote in collaboration with Amit Sawant, PhD, at the University of Maryland, Baltimore, and radiation oncologists will be able to predict movement in lung tumors as the patient breathes in real-time with a 3D motion model.
New research at the Scientific Computing and Imaging Institute at the University of Utah, helps to precisely target tumors in lung cancer patients using artificial intelligence. Employing algorithms developed by Sarang Joshi, DSc, professor of Biomedical Engineering and graduate student Markus Foote in collaboration with Amit Sawant, PhD, at the University of Maryland, Baltimore, and radiation oncologists will be able to predict movement in lung tumors as the patient breathes in real-time with a 3D motion model.
SCI Head Model Now Available
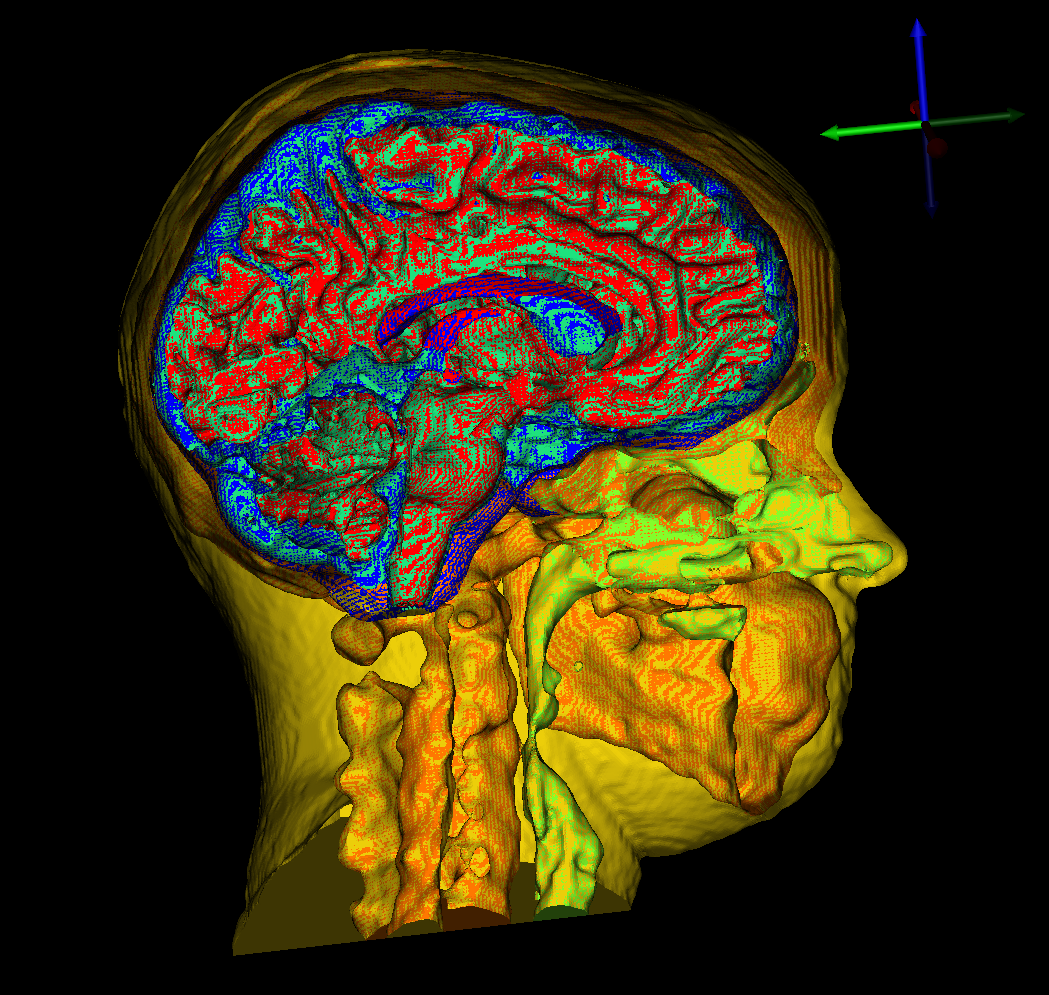 The SCI Institute is excited to announce the release of the SCI Institute's female head and brain model data repository and pipeline.
The SCI Institute is excited to announce the release of the SCI Institute's female head and brain model data repository and pipeline. The dataset and corresponding publication can be found here.
What you can’t see can hurt you
 You can't see nasty microscopic air pollutants in your home, but what if you could?
You can't see nasty microscopic air pollutants in your home, but what if you could?Engineers from the University of Utah's School of Computing conducted a study to determine if homeowners change the way they live if they could visualize the air quality in their house. It turns out, their behavior changes a lot.
3D Virtual Simulation Gets to the ‘Heart’ of Irregular Heartbeats
A 3-D virtual heart. Credit: Johns Hopkins University
Contributions to Computer Graphics

By Paul Gabrielsen, science writer, University of Utah Communications
On Monday, August 13, the computer graphics world honored 52 people as inaugural members of the SIGGRAPH Academy. Each played a role in bringing graphics from their beginnings as arrangements of glowing lines to the immersive 3D worlds that moviemakers, game designers and others build today. Seven of those 52 were alumni of the University of Utah. And one was the "father of computer graphics," former professor Ivan Sutherland.
At the Hayden Planetarium, a Joyride Across the Cosmos
Story appears at the New York Times
 Carter Emmart used to want to go to space. Now he does, all the time — but virtually. And he likes to share.
Carter Emmart used to want to go to space. Now he does, all the time — but virtually. And he likes to share.On a recent afternoon, he took a visitor at the Hayden Planetarium, where he works, on a kind of joyride across the known universe. The lights dim. The inverted bowl of the planetarium’s screen comes to life. But the familiar, insectlike projector that displays the stars and constellations is stowed under the floor. Instead, projectors are reconstructing images onto the half dome from a desktop computer.
So there’s Mars, filling the screen, its reddish desert revealed as a landscape of mountains and valleys that make the Grand Canyon look like a puny arroyo. Flying around, we take in the sights, the surroundings uncannily close and detailed, so that boulders only a few feet across can be discerned.
Machine learning could improve how doctors diagnose heart attacks
 You're working in your house, going about your normal routine when suddenly the pain hits. Your chest starts to throb and your left arm begins to ache. Without hesitation, you rush to the hospital, dreading your worst fear has become a reality — you are having a heart attack. Upon arrival, physicians, nurses, and other medical staff begin frantically testing, probing, and prodding nearly every part of your body. They run more tests than you can keep track of and begin shouting orders for new tests and other members of the team. The emergency physician is carefully watching the monitors hooked up by your bedside, puzzled by the results they are seeing. They turn to consult a cardiologist expert on the signals your heart is emitting. But instead of a person, they turn to a computer.
You're working in your house, going about your normal routine when suddenly the pain hits. Your chest starts to throb and your left arm begins to ache. Without hesitation, you rush to the hospital, dreading your worst fear has become a reality — you are having a heart attack. Upon arrival, physicians, nurses, and other medical staff begin frantically testing, probing, and prodding nearly every part of your body. They run more tests than you can keep track of and begin shouting orders for new tests and other members of the team. The emergency physician is carefully watching the monitors hooked up by your bedside, puzzled by the results they are seeing. They turn to consult a cardiologist expert on the signals your heart is emitting. But instead of a person, they turn to a computer.
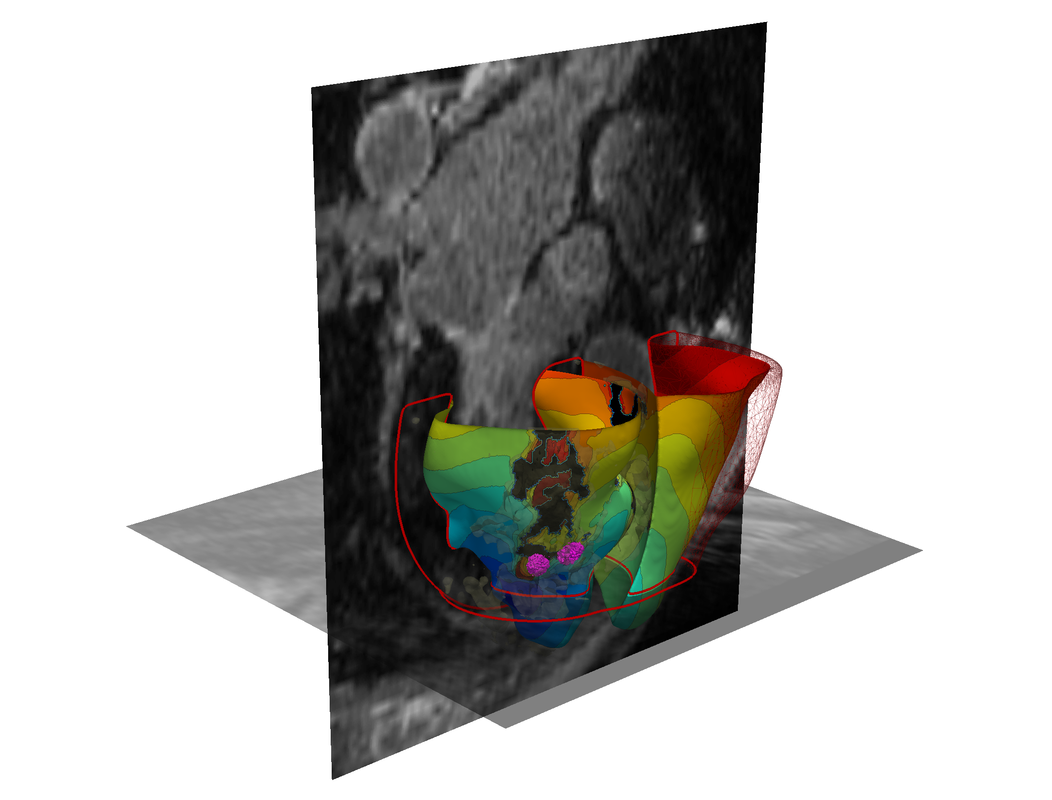
Targeting Tumors More Accurately
 New research being conducted at the Scientific Computing and Imaging Institute holds the potential to increase the accuracy of targeted treatments for tumors in the lungs. Currently, motion caused by the patient's breathing introduces motion artifacts when imaging lung tumors. The inherent breathing motion also limits the precise targeting of radiation therapy for treating lung cancer.
New research being conducted at the Scientific Computing and Imaging Institute holds the potential to increase the accuracy of targeted treatments for tumors in the lungs. Currently, motion caused by the patient's breathing introduces motion artifacts when imaging lung tumors. The inherent breathing motion also limits the precise targeting of radiation therapy for treating lung cancer.
Nothing is Certain
 "The only certainty...," it is said, "is that nothing is certain."
"The only certainty...," it is said, "is that nothing is certain."
And so it goes with computational forecasts of important events such as weather, finance, and climate. Among all of this uncertainty, however, there are patterns, likelihoods, and rarities that inform important decisions that may affect billions of dollars in resources and thousands, or even millions, of lives. In the hurricane season on the eastern U.S., computational forecasting plays a central role in critical decisions that can determine allocations of emergency resources and the movements of people. The uncertainty and accuracy of these forecasts is an important part in making effective use of these sophisticated tools.
Need for Speed
 Whether it’s coming up with the best design for a Formula 1 race car or understanding the effects of atrial fibrillation on the heart, developing the right simulation model for research sometimes involves equal parts applied math, engineering and computer science.
Whether it’s coming up with the best design for a Formula 1 race car or understanding the effects of atrial fibrillation on the heart, developing the right simulation model for research sometimes involves equal parts applied math, engineering and computer science.University of Utah School of Computing professor Mike Kirby sees himself as the person who connects these disciplines so he can take trailblazing ideas and help create better simulation software to aid researchers.
A Calming Effect
 For many who suffer from debilitating neurological disorders such as Parkinson’s Disease, the constant muscular tremors are an unbearable symptom. Just drinking from a cup can be an overwhelming challenge.
For many who suffer from debilitating neurological disorders such as Parkinson’s Disease, the constant muscular tremors are an unbearable symptom. Just drinking from a cup can be an overwhelming challenge.When medication doesn’t work, brain surgery to destroy certain cells can be risky, and the results are irreversible. But there has been an emerging third option — deep brain stimulation (DBS), a therapy in which electrodes are implanted in the patient’s brain that deliver continuous electrical pulses to control motor function.
University of Utah bioengineering associate professor Christopher Butson has been researching ways to improve DBS systems to make them more effective and convenient for patients who wear them. He believes an answer lies in mobile tablets and smartphones.
Gigapixel image analysis on the fly
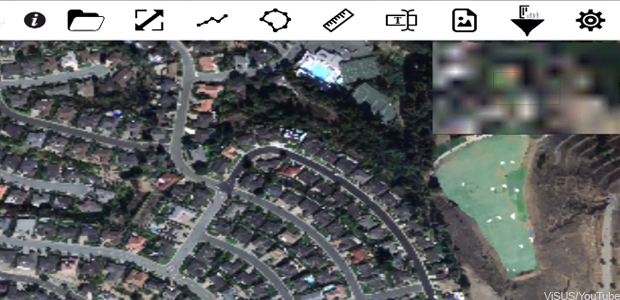 Most farmers probably never thought they'd be in the market for a way to process huge digital images more quickly -- until, that is, inexpensive drones with high-resolution cameras gave them access to images they could use to micromanage irrigation and to detect the growth of crop-threatening diseases.
Most farmers probably never thought they'd be in the market for a way to process huge digital images more quickly -- until, that is, inexpensive drones with high-resolution cameras gave them access to images they could use to micromanage irrigation and to detect the growth of crop-threatening diseases.
2016 Image-Based Biomedical Modeling (IBBM) summer course
 The Image-Based Biomedical Modeling (IBBM) summer course was held for the third time in 2016, from July 11-21, 2016 in Newpark Resort, Park City (Utah). The two-week course format included the following activities:
The Image-Based Biomedical Modeling (IBBM) summer course was held for the third time in 2016, from July 11-21, 2016 in Newpark Resort, Park City (Utah). The two-week course format included the following activities:- Didactic lecture sessions given by the three PIs (Rob MacLeod, Ross Whitaker and Jeff Weiss) as well as three invited instructors (Miriah Meyer, Steve Maas and Gerard Ateshian) experts in their fields
- Laboratory exercises lead by a group of teaching assistants and developers,
- Discussion session time for student-instructor interaction,
- Visit to the experimental and computational laboratory facilities at the University of Utah, College of Engineering to give the participants an overview of the general academic background and research projects performed at the university,
- Four Keynotes Lectures from leaders in the field,
- Mentoring lectures on grant writing, responsible conduct of research, and simulation study design.
Introduction to Image-Based Modeling
 |
 |
 |
 |
 |
 |
Image Analysis Tools for Understanding Connective Tissue Structure
 Sponsored by the Burton Foundation
Sponsored by the Burton Foundation
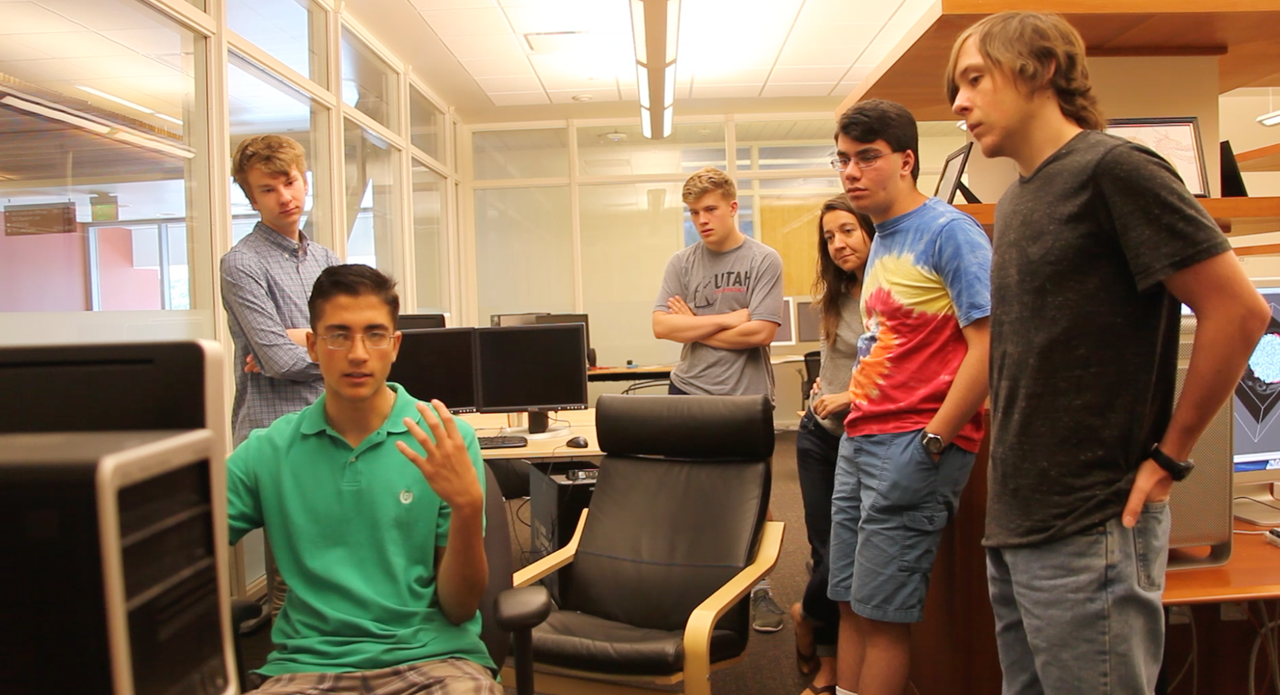
This summer, two Salt Lake area high school students from Copper Hills High School came to the University of Utah to participate in a hands-on research experience. The students learned how image analysis tools help biomechanics researchers understand the effects of structural features of musculoskeletal tissues (e.g. tendons, ligaments, and articular cartilage) on the functional behavior of these tissues.Many musculoskeletal tissue injuries and diseases exhibit altered macroscopic and microscopic tissue structure. The Musculoskeletal Research Laboratories, a research center of the Scientific Computing and Imaging Institute, uses engineered tissue materials to study the effect of these structural changes on tissue behavior. Researchers use many image acquisition techniques to characterize the structure of native and engineered tissues, including optical microscopy, x-ray computed tomography (CT), and electron microscopy. Image analysis tools allow efficient detection and quantification of structural features from these images.
Visualizing Hurricanes
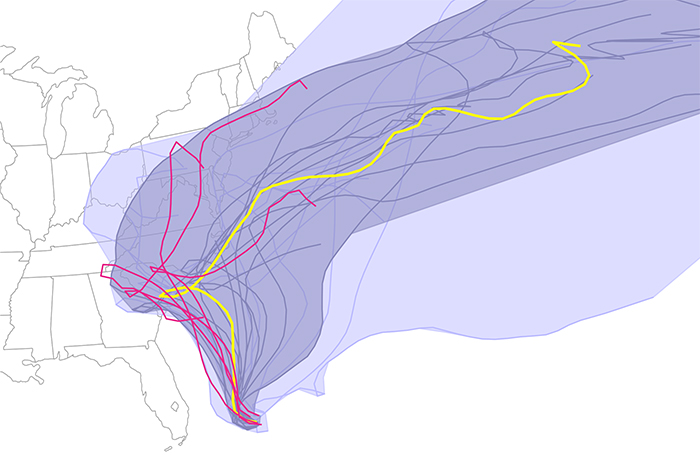 |
| The results denoted possible predicted paths, based upon different models and/or conditions Joaquin might take as of Friday October 2, 2015. Using their Curve Boxplot analysis and visualization method, they show the median hurricane path and the 50 percent band (dark region) — denoting the spatial swath in which 50 percent of the predicted hurricane tracks lie. The light band denotes nearly 100 percent of the possible paths predicted. Red denotes outliers — those hurricane paths flagged as unlikely in reference to all other members of the ensemble. |
2015 Image-Based Biomedical Modeling (IBBM) summer course
 The two-week course included the following activities:
The two-week course included the following activities:- Didactic lecture sessions given by the three PIs as well as four invited instructors and experts in their fields .
- Laboratory exercises led by a group of 10 teaching assistants and developers.
- Discussion session time for student-instructor interaction.
- A visit to the experimental and computational laboratory facilities at the University of Utah, College of Engineering to give the participants an overview of the general academic background and research projects performed at the university.
- Mentoring lectures on grant writing, responsible conduct of research, and simulation study design.
2014 Summer Course on Image-based Biomedical Modeling (IBBM)
The Image-Based Biomedical Modeling (IBBM) summer course was held from July 14 to July 24 in the Newpark Hotel, Park City, Utah.
The two-week summer course hosted 39 participants this year: 31 graduate students, 1 MD/PhD student, 2 postdoctoral fellows, 3 junior faculty, and 2 developers from a research laboratory / industry. Participants came from 24 institutions, including 4 from universities in Belgium and England. After the first week of common classes, participants were divided into two tracks: Bioelectricity (10 participants) and biomechanics (29 participants).
 IBBM is a dedicated two-week course in the area of image-based modeling and simulation applied to bioelectricity and biomechanics, providing participants with training in the numerical methods, image analysis, visualization, and computational tools necessary to carry out end-to-end, image-based, subject-specific simulations in either bioelectricity or orthopedic biomechanics. The course focuses on using freely available, open-source software developed under the research of the CIBC (P41 GM103545) and FEBio suite (RO1 GM083925). Students use this software to learn and apply the complete dataflow pipeline to particular sets of data with specific goals.
IBBM is a dedicated two-week course in the area of image-based modeling and simulation applied to bioelectricity and biomechanics, providing participants with training in the numerical methods, image analysis, visualization, and computational tools necessary to carry out end-to-end, image-based, subject-specific simulations in either bioelectricity or orthopedic biomechanics. The course focuses on using freely available, open-source software developed under the research of the CIBC (P41 GM103545) and FEBio suite (RO1 GM083925). Students use this software to learn and apply the complete dataflow pipeline to particular sets of data with specific goals.
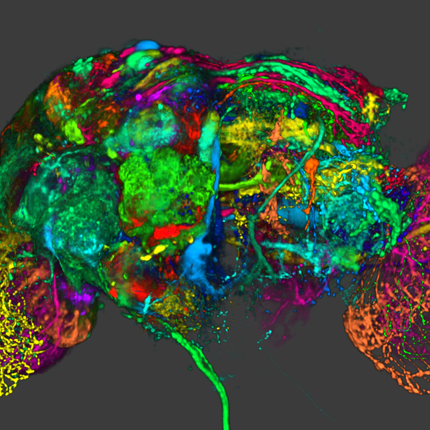
Understanding the Morphology of Brain Disorders
Advances in medical imaging devices, such as magnetic resonance imaging (MRI), have led to our ability to acquire detailed information about the living human brain, including its anatomical structure, function, and connectivity. However, making sense of this complex data is a difficult task, especially in large imaging studies that may include hundreds or even thousands of participants. This is where computer science can play an important role. Image analysis algorithms can automatically quantify properties of the brain, such as the size of brain structures, or the functional activity in different brain regions. This provides neuroscience researchers with insights into how the brain functions and what abnormalities are present in diseased brains.
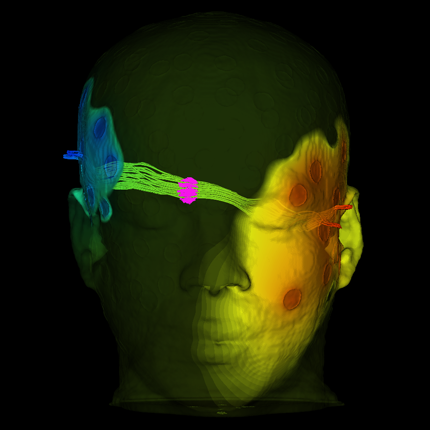
Neuro Stimulation
 |
| Transcranial Magnetic Stimulation (TMS) of the human motor cortex. |