News
NeuroImage Journal Cover Features SCIRun Renderings
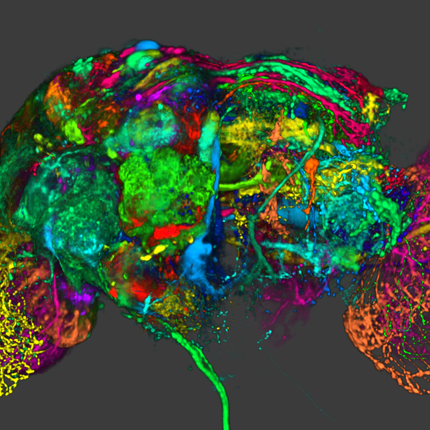
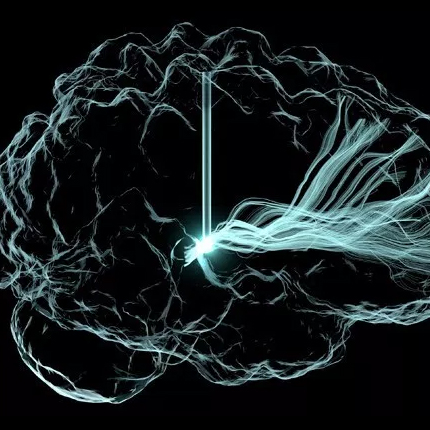
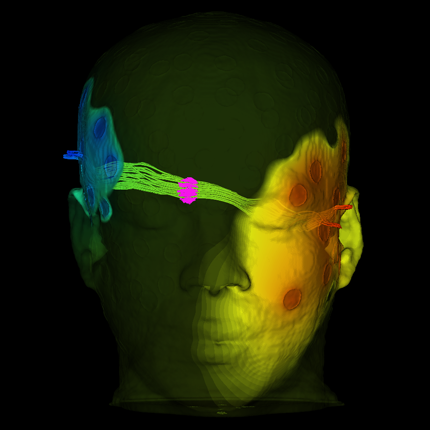
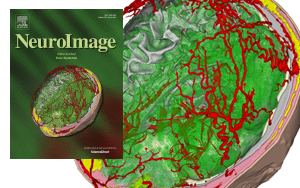 The publication and cover image is the result of a close collaboration between the University of Freiburg (Lukas Fiederer, Tonio Ball and others) in Germany and the NIH-funded Center for Integrated Biomedical Computing (CIBC, Moritz Dannhauer) at the SCI institute (Johannes Vorwerk). The cover of NeuroImage' March (128) issue illustrates different tissues in a model of the human head that are known to have distinct electrical properties. In this study, we simulated the electrical effect of blood vessels on current flow originating from active brain regions as monitored by scalp electrodes (encephalography, EEG). Since the understanding of EEG measurements matters in many clinical applications (e.g., epilepsy), SCIRun software offers a set of tools to investigate their underlying electrical generators. Recently, in the new version 5.0 of SCIRun additional capabilities (BrainStimulator) have been added to simulate the current flow from external stimulation devices targeting brain regions of potential EEG generators.
The publication and cover image is the result of a close collaboration between the University of Freiburg (Lukas Fiederer, Tonio Ball and others) in Germany and the NIH-funded Center for Integrated Biomedical Computing (CIBC, Moritz Dannhauer) at the SCI institute (Johannes Vorwerk). The cover of NeuroImage' March (128) issue illustrates different tissues in a model of the human head that are known to have distinct electrical properties. In this study, we simulated the electrical effect of blood vessels on current flow originating from active brain regions as monitored by scalp electrodes (encephalography, EEG). Since the understanding of EEG measurements matters in many clinical applications (e.g., epilepsy), SCIRun software offers a set of tools to investigate their underlying electrical generators. Recently, in the new version 5.0 of SCIRun additional capabilities (BrainStimulator) have been added to simulate the current flow from external stimulation devices targeting brain regions of potential EEG generators.
Best Paper Award at PacificVis 2016
 Congratulations Bei and Valerio and SCI alumni: Paul Rosen, Guoning Chen and Harsh Bhatia who were awarded Best Paper at IEEE PacificVis 2016.
Congratulations Bei and Valerio and SCI alumni: Paul Rosen, Guoning Chen and Harsh Bhatia who were awarded Best Paper at IEEE PacificVis 2016.Critical Point Cancellation in 3D Vector Fields: Robustness and Discussion Primoz Skraba (Jozef Stefan Institute), Paul Rosen (University of South Florida), Bei Wang (University of Utah), Guoning Chen (University of Houston), Harsh Bhatia (Lawrence Livermore National Laboratory), Valerio Pascucci (University of Utah)
Science-ArtQuiz
 The SCI Institute recently contributed to the Science-ArtQuiz with the Leonardo. Can you distinguish Science from Art???
The SCI Institute recently contributed to the Science-ArtQuiz with the Leonardo. Can you distinguish Science from Art???Paintbrush or microscope? Einstein or Picasso? Is there any difference? Challenge your inner genius with this unique, mind bending science vs art quiz. Can you get a perfect score?
Take the quiz at http://www.theleonardo.org/science-artquiz/
Mike Kirby among coauthors of ISC 2016 award winning paper

Congratulations to James King, Thomas Gilray, Robert M. Kirby and Matthew Might, whose paper: Dynamic Sparse-Matrix Allocation on GPUs, won the International SuperComputing PRACE-ISC Award 2016
NSF Graduate Fellowships Awarded to Two SCI Students
 Congratulations to Danielle Pruss and Daria Anderson on receiving NSF Graduate Fellowships. Danielle is a researcher in the Vis Design Lab under Dr. Miriah Meyer, and Daria is working on issues with deep brain stimulation under Dr. Chris Butson.
Congratulations to Danielle Pruss and Daria Anderson on receiving NSF Graduate Fellowships. Danielle is a researcher in the Vis Design Lab under Dr. Miriah Meyer, and Daria is working on issues with deep brain stimulation under Dr. Chris Butson.
SCI Research Highlighted in Argonne's 10 Year Celebration
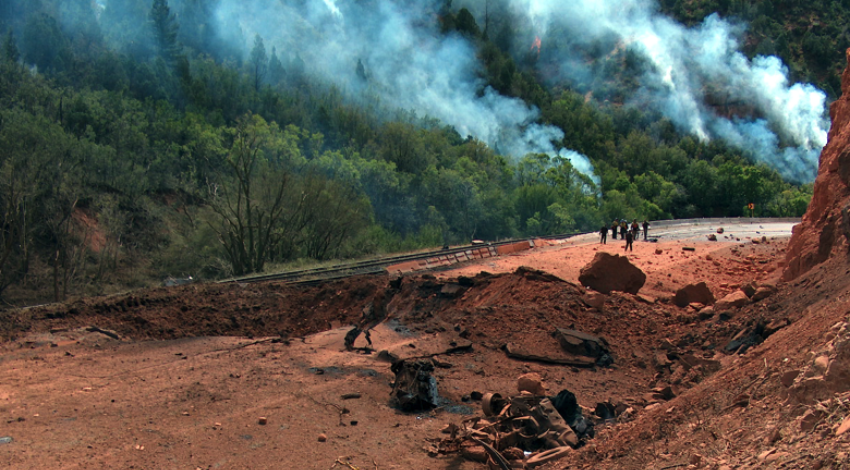 Modeling detonations to transport explosives safely
Modeling detonations to transport explosives safelyResearchers modeled a 2005 explosion that left a 30-by-70-foot crater in a Utah highway, capturing the physics that made the truck's cargo explode more violently than it should have. With such simulations, we can design safer transport for explosives. Led by Martin Berzins, University of Utah
You can see the 10 highlights at https://www.alcf.anl.gov/articles/10-science-highlights-celebrating-10-years-argonne-leadership-computing-facility
Chris Johnson Re-Elected to CRA Board of Directors
 Congratulations to Chris Johnson on being re-elected for another three years to the Computing Research Association board of directors.
Congratulations to Chris Johnson on being re-elected for another three years to the Computing Research Association board of directors. You can read the full release here.
Multiple Awards Received at IVAPP 2016
 Congratulations to Hao Nguyen for winning both best paper and best PhD progect at the International Conference on Information Visualization Theory and Applications (IVAPP).
Congratulations to Hao Nguyen for winning both best paper and best PhD progect at the International Conference on Information Visualization Theory and Applications (IVAPP).Best Paper: H. Nguyen, P. Rosen, Improved identification of data correlations through correlation coordinate plots, Intl. Conf. on Information Visualization Theory and Applications (IVAPP), 2016.
Best PhD Project: H. Nguyen, P. Rosen, Data Scalable Approach for Identifying Correlation in Large and Muti-Dimensional Data, Intl. Conf. on Information Visualization Theory and Applications Doctoral Consortium (IVAPP), 2016.
Seg3D 2.3.0 Released
 We are pleased to announce Seg3D version 2.3.0.
We are pleased to announce Seg3D version 2.3.0.The new binary installers and sources can be downloaded from:
https://github.com/SCIInstitute/Seg3D/releases
Behind Each Breath, an Underappreciated Muscle
 CIBC collaborator Gabrielle Kardon's research has been showcased by New York Times science writer Carl Zimmer.
CIBC collaborator Gabrielle Kardon's research has been showcased by New York Times science writer Carl Zimmer. You can read the full article on their website.
Winner of the 2015 Teapot Rendering Competition
This image is not strictly photorealistic. Laura rendered out the image into two main layers, one for specular highlights and caustics, and one for diffuse lighting. She also has a mask for just the teapots, and one for the ground. Laura was able to split up the image and adjust the brightness of each part independently. Here she decided to include some diffuse reflection/refraction for the teapots, but mostly removed the diffuse lighting from the ground. This gave an extra glow to the teapots and also emphasized the caustics. 100,000 samples per pixel.